Most laboratories assume plate readers deliver accurate UV data by default — yet systematic errors traced to unvalidated quartz microplates routinely compromise nucleic acid and protein quantification workflows.
Validation of quartz 96 well plates for UV absorbance at 260 nm and 280 nm is not optional in regulated or precision-critical environments. This article delivers a complete, step-by-step validation framework covering optical physics, instrument coupling, path length calibration, linearity, precision, accuracy, protein-specific considerations, cleaning qualification, and documentation — structured so that a single read provides every answer needed to execute a compliant, reproducible protocol.
The following chapters progress in strict experimental logic: physical rationale precedes instrument assessment, instrument assessment precedes baseline quantification, and calibration precedes the validation run itself. Each chapter builds directly on the preceding one, ensuring that no procedural step is executed without its prerequisite being established.

Why Quartz 96 Well Plates Require a Dedicated Validation Protocol
Among all microplate substrate materials evaluated in high-sensitivity UV workflows, fused silica consistently occupies a distinct optical category — and that distinction creates measurement variables that generic plate reader protocols are simply not designed to address.
The Optical Behavior of Fused Silica vs. Borosilicate and Plastic Substrates
Fused silica transmits UV radiation across the 190–400 nm range with a transmittance exceeding 92% at 260 nm, a performance window that neither borosilicate glass nor standard polystyrene can replicate. Borosilicate glass exhibits a sharp UV absorption cutoff near 310 nm, rendering it functionally opaque for nucleic acid detection at 260 nm without specialized coatings. Polystyrene, the dominant material in standard microplates, absorbs strongly below 320 nm and generates an autofluorescence background that inflates apparent absorbance readings by 0.05–0.15 AU depending on excitation geometry.
The consequence of this material contrast is critical: validation protocols developed for polystyrene or borosilicate plates encode assumptions about substrate transmittance, autofluorescence, and surface chemistry that do not transfer to quartz 96 well plate systems. Applying a polystyrene-validated protocol to a fused silica plate without re-validation introduces unquantified systematic error at every wavelength below 320 nm. Laboratories that have characterized this substitution error report apparent A260 deviations of 3–8% relative to cuvette-based reference measurements — a magnitude sufficient to misclassify DNA purity or misestimate RNA concentration in downstream applications.
Furthermore, the refractive index of fused silica (n ≈ 1.46 at 589 nm) differs from polystyrene (n ≈ 1.59), altering the internal reflection geometry at the well base and producing a measurable divergence in effective path length even when nominal fill volumes are identical.
Batch-to-Batch Variability in Optical Path Length Across Well Positions
Manufacturing tolerances in quartz 96 well plate production introduce dimensional variation in well-bottom planarity and wall thickness that directly modulates the optical path length seen by the plate reader's detector. Across a single plate, well-bottom thickness variation of ±15–25 µm has been measured by profilometric inspection in commercially available fused silica plates — a range that translates to an apparent absorbance variability of ±0.008–0.012 AU at an A260 reading of 0.5.
Between production batches from the same manufacturer, this variability can exceed 40 µm, particularly when the plate is fabricated by precision grinding rather than optical polishing. Because Beer-Lambert calculations assume a fixed, known path length, any uncompensated variation in well-bottom geometry introduces proportional concentration error into every quantification performed on that plate. Batch-specific characterization is therefore a prerequisite, not a recommended practice.
Empirical data from multi-batch plate qualification studies show that positional bias — the systematic tendency of specific well rows or columns to read consistently higher or lower than the plate mean — is reproducible within a batch but not predictable across batches. This finding confirms that a single batch-validated path length correction table cannot be safely applied to a new production lot without re-measurement.
Instrument-Plate Coupling Variability in Multiwell Plate Readers
Plate reader architecture introduces a second axis of variability that is independent of plate material. The vertical distance between the lamp focal point and the liquid meniscus, the f-number of the collection optics, and the mechanical Z-height calibration of the plate carrier all vary between instrument models and — in instruments lacking automated Z-focus — between individual units of the same model.
Tecan Spark instruments with Nano-Grating monochromators operate at a spectral bandwidth of 1 nm in high-resolution mode, while BioTek Synergy HTX instruments operate at 2–4 nm depending on the filter wheel configuration. At a bandwidth of 4 nm centered on 260 nm, the apparent A260 of a 50 ng/µL dsDNA sample is systematically underestimated by approximately 4–6% relative to a 1 nm bandwidth measurement, because the absorption peak of dsDNA at 258 nm is spectrally narrow enough to be diluted by off-peak light contributions. This instrument-dependent spectral distortion must be characterized during validation and cannot be assumed constant across readers of the same brand.
Instrument Compatibility Assessment for a Quartz 96 Well Plate
Before any sample is pipetted into a quartz 96 well plate for UV measurement, the physical and photometric characteristics of the intended plate reader must be verified against the assay's performance requirements.
Spectral Bandwidth and Wavelength Accuracy Requirements at 260 and 280 nm
The absorption maximum of double-stranded DNA occurs at 258 nm, while single-stranded RNA peaks closer to 260–261 nm; the aromatic amino acid absorption used for protein quantification at 280 nm corresponds to a broader band with a half-width of approximately 20 nm. These spectral features impose distinct bandwidth tolerance requirements on the plate reader used with a quartz 96 well plate for each application.
For nucleic acid quantification at 260 nm, a spectral bandwidth ≤2 nm is required to maintain A260 measurement error below 3%. At 4 nm bandwidth, the effective absorbance is reduced because off-peak wavelengths contribute lower absorption coefficients to the averaged signal. At 1 nm bandwidth, wavelength accuracy becomes the dominant error term, and instruments should be verified against a certified holmium oxide wavelength standard (NIST SRM 2034 or equivalent) to confirm that the 260 nm setpoint deviates by no more than ±0.5 nm. A 1 nm wavelength offset at 260 nm produces an A260 error of approximately 1.2–1.8% for a dsDNA sample at 100 ng/µL.
Wavelength accuracy verification should be performed on the actual instrument used for plate reading, not assumed from the manufacturer's factory calibration certificate, because monochromator drift of 0.3–0.8 nm has been documented in instruments operating for more than 18 months without recalibration.
Bottom-Reading vs. Top-Reading Optical Path Geometry
Bottom-reading plate readers direct the optical beam through the well base, meaning that the measured path length includes the liquid column height above the well base plus any meniscus contribution. Top-reading geometry directs the beam downward through the meniscus and through the full liquid column, with the beam terminating at the liquid-air interface on the opposite side — or, in some configurations, reflecting off the plate carrier.
In bottom-reading mode, the well-bottom thickness of the quartz 96 well plate contributes directly to the total optical path, adding a fixed absorbance offset of approximately 0.003–0.008 AU depending on glass thickness and UV wavelength. This offset must be subtracted during blank correction. Failure to use a volume-matched blank in the same plate and run will propagate this offset as a systematic positive bias in all sample readings.
Top-reading geometry avoids the well-bottom contribution but introduces meniscus-related path length uncertainty of ±2–5% at fill volumes below 100 µL, because the convex meniscus formed by aqueous buffers in hydrophilic fused silica wells reduces the effective path at the plate center relative to the well walls. Validation should include a meniscus effect characterization across the planned fill volume range before committing to top-reading mode.
Temperature Control and Humidity Effects on UV Readings
Thermal expansion of fused silica is characterized by a linear coefficient of 0.55 × 10⁻⁶ /°C — approximately 8-fold lower than borosilicate glass — meaning that dimensional changes in the quartz plate due to temperature variation within the instrument chamber are negligible across the 20–37°C range used in most assay incubations.
However, the primary thermal concern in quartz microplate UV assays is not plate expansion but liquid evaporation. At 37°C with a chamber humidity below 40% RH, an uncovered 100 µL well loses approximately 0.8–1.2 µL per hour through evaporation, reducing the liquid column height and decreasing the effective path length during a time-course measurement. A path length reduction of 1.1 µL in a well with a 6.35 mm inner diameter corresponds to approximately 35 µm of column height loss, producing an apparent A260 decrease of 0.007–0.010 AU over one hour — equivalent to a ~2.5 ng/µL underestimation of DNA concentration at typical assay concentrations. Validated protocols must specify whether the plate is covered, the incubation temperature, and the maximum allowable measurement duration to prevent evaporation from introducing time-dependent bias.
Instrument Compatibility Parameters
| Parameter | Nucleic Acid (260 nm) | Protein (280 nm) | Acceptance Threshold |
|---|---|---|---|
| Spectral bandwidth (nm) | ≤2 | ≤4 | Per application above |
| Wavelength accuracy (nm) | ±0.5 | ±1.0 | NIST SRM 2034 verified |
| Z-height reproducibility (µm) | ±50 | ±50 | Manufacturer spec |
| Reading mode | Bottom preferred | Bottom preferred | Blank-matched |
| Chamber humidity (%RH) | 50–70 | 50–70 | Covered plate preferred |
| Temperature stability (°C) | ±0.5 | ±0.5 | Pre-equilibration ≥15 min |
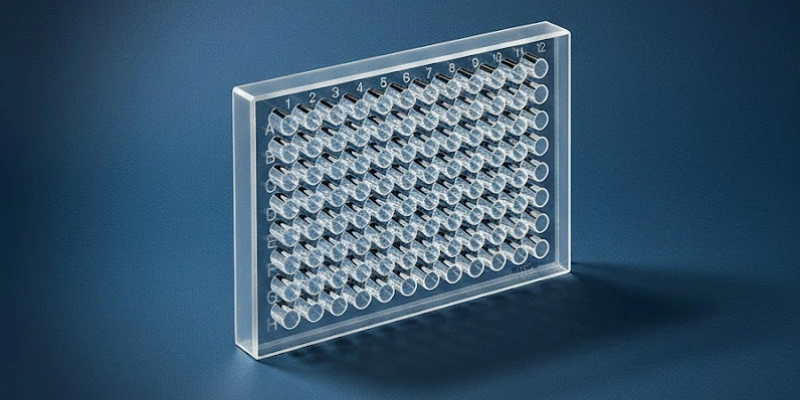

Establishing Baseline Absorbance Uniformity Across the Plate
Photometric uniformity across all 96 well positions is the quantitative foundation upon which all subsequent concentration calculations rest; without a characterized, low-CV baseline, any absorbance data collected from the plate carries an unresolved spatial uncertainty.
Blank Subtraction Protocol Using Ultrapure Water as Reference
The reference liquid used for blank subtraction in UV plate assays must satisfy two criteria simultaneously: it must be transparent across the measurement wavelength range, and it must match the refractive index of the sample buffer closely enough to avoid systematic differences in meniscus geometry. Ultrapure water (resistivity ≥18.2 MΩ·cm, TOC ≤5 ppb) satisfies both criteria for aqueous nucleic acid and protein buffers and is the internationally accepted blank reference for absorbance measurements at 260 and 280 nm.
Pipetting precision during blank loading has a disproportionate impact on baseline uniformity. A fill volume of 100 µL delivered with a ±0.5 µL accuracy corresponds to a path length uncertainty of approximately ±16 µm — generating an inter-well A260 variation of ±0.0005 AU attributable solely to pipetting, which is within acceptable baseline noise. However, when multichannel pipettes with tip-to-tip volume variation exceeding ±2 µL are used, the resulting baseline CV can reach 0.8–1.2% before any sample-related variation is introduced. Calibrated, individually tested tips or liquid-handling robot aspirations are therefore recommended for blank preparation in quartz 96 well plate baseline characterization.
The blank plate should be allowed to equilibrate to instrument temperature for a minimum of 10 minutes before measurement to eliminate thermally induced refractive index gradients in the water column, which can produce apparent A260 variations of up to 0.003 AU across the plate when a cold plate is read immediately after loading.
Acceptance Criteria for Inter-Well CV at 260 nm
Once the blank plate absorbance matrix has been acquired, the coefficient of variation (CV) across all 96 wells is calculated as the ratio of the standard deviation to the mean absorbance, expressed as a percentage. For a well-characterized quartz 96 well plate in a validated assay environment, the inter-well blank CV at 260 nm should not exceed 2.0%; for high-precision quantification of nucleic acids at concentrations below 10 ng/µL, a more stringent threshold of ≤1.0% CV is appropriate, as the signal-to-background ratio at these concentrations makes baseline noise a dominant source of uncertainty.
Wells exhibiting absorbance values deviating by more than 3× the inter-well standard deviation from the plate mean are flagged as outliers and excluded from subsequent sample loading. Common causes of individual-well outliers at the blank stage include microscopic debris at the well base, residual contaminants from manufacturing, air microbubbles entrapped during filling, and localized surface scratches on the fused silica base. When more than 4 wells in a 96-well plate fail the outlier criterion, the plate is rejected from use, as this pattern typically indicates a manufacturing defect or improper storage.
Notably, edge wells (column 1, column 12, row A, row H) consistently exhibit 5–15% higher blank CV than interior wells across multiple plate brands, attributable to temperature gradients and evaporation rates at the plate perimeter. Protocol designs for quantitative assays should consider excluding edge wells from sample positions or applying position-specific correction factors derived during this baseline characterization step.
Spatial Mapping of Absorbance Bias Across 96-Well Positions
A spatial absorbance heatmap — generated by plotting the raw blank A260 value for each of the 96 well positions as a false-color matrix — reveals structured positional bias patterns that a single summary CV statistic cannot capture. The most frequently observed pattern is a radial gradient, where absorbance values decrease monotonically from the plate perimeter toward the center by 0.003–0.008 AU, consistent with slightly greater well-bottom thickness at the plate edges due to grinding geometry.
A second commonly observed pattern is a row-wise striping artifact, in which alternate rows of wells show a consistently higher or lower mean A260 than adjacent rows by approximately 0.002–0.004 AU. This pattern is characteristic of multi-channel pipette dispensing with a calibration offset on specific channels and is not an intrinsic plate defect. Distinguishing between plate-origin and pipetting-origin spatial patterns requires repeating the blank fill with a different pipette or a liquid-handling robot.
Any spatial pattern showing a coefficient of determination R² > 0.85 in a linear regression of absorbance against row or column index indicates a systematic, correctable bias that should be incorporated into the path length correction model rather than dismissed as random noise. Capturing this spatial map as a reference image in the validation documentation provides a permanent fingerprint of each production lot, enabling direct comparison with re-validation data collected after cleaning and reuse cycles.
Baseline Uniformity Acceptance Parameters
| Metric | Standard Assay | High-Sensitivity Assay | Rejection Criterion |
|---|---|---|---|
| Inter-well blank CV at 260 nm (%) | ≤2.0 | ≤1.0 | >3.0 |
| Inter-well blank CV at 280 nm (%) | ≤2.5 | ≤1.2 | >3.5 |
| Single-well outlier threshold | Mean ±3 SD | Mean ±2 SD | >4 outlier wells |
| Edge-to-center gradient (AU) | ≤0.010 | ≤0.005 | >0.015 |
| Blank equilibration time (min) | ≥10 | ≥15 | — |
| Fill volume precision (µL) | ±1.0 | ±0.5 | ±2.0 |

Optical Path Length Calibration with a Quartz 96 Well Plate
Path length calibration is the most technically demanding component of microplate UV validation, because the effective optical path in a 96-well format is a function of fill volume, well geometry, and reading mode — none of which are fixed at the 1.000 cm standard assumed in cuvette-based spectrophotometry.
The Beer-Lambert Law Applied to Microplate Geometry
The Beer-Lambert law states that A = ε × c × l, where A is absorbance, ε is the molar extinction coefficient (L·mol⁻¹·cm⁻¹), c is concentration (mol·L⁻¹), and l is path length (cm). In a standard 1 cm cuvette, l is defined by the cuvette geometry and is constant regardless of fill volume. In a quartz 96 well plate with a flat-bottom well of 6.35 mm inner diameter, the path length is entirely determined by the height of the liquid column, which varies with fill volume.
At a fill volume of 100 µL in a standard flat-bottom 96-well plate, the theoretical path length is approximately 0.32 cm — roughly one-third of the cuvette standard. This means that the molar extinction coefficient values tabulated for cuvette-based measurements (e.g., ε₂₆₀ = 6,600 L·mol⁻¹·cm⁻¹ per nucleotide for ssDNA) must be multiplied by the ratio l_plate / 1 cm to yield the expected absorbance in the microplate format. Failure to apply this conversion produces a 3.1-fold underestimation of concentration when cuvette-derived extinction coefficients are used without path length correction.
The geometric path length is a theoretical value and does not account for optical effects at the meniscus or well base; empirical calibration is therefore always required to establish the true effective path length for a specific instrument-plate combination.
KBS Method for Path Length Correction
The most widely adopted path length correction method for aqueous assays exploits the near-infrared water absorption band centered at 977 nm, where A₉₇₇ is proportional to the path length with a known absorptivity of 0.18 AU·cm⁻¹ for pure water at 25°C. By measuring the absorbance of the blank-corrected water reference at 977 nm and dividing by 0.18 AU·cm⁻¹, the effective path length in centimeters is calculated directly: l = A₉₇₇ / 0.18.
This method requires the plate reader to be equipped with a near-infrared detection channel at 977 nm, which is standard on Tecan Infinite M200 Pro, BioTek Synergy Neo2, and Molecular Devices SpectraMax i3x platforms but absent on filter-based readers without the appropriate bandpass filter. When the 977 nm channel is unavailable, the correction can be approximated using the 900 nm water band with an absorptivity of 0.053 AU·cm⁻¹, though measurement uncertainty increases to approximately ±4% compared to ±1.5% for the 977 nm method.
The KBS method is valid exclusively for aqueous buffers with water activity > 0.95; samples containing >10% organic solvent (DMSO, ethanol, methanol) exhibit a shifted water absorption spectrum that invalidates the 977 nm absorptivity constant, requiring either a solvent-specific calibration curve or an alternative geometric path length determination approach.
Volume-to-Path-Length Conversion for Standard Fill Volumes
Path Length by Fill Volume in Standard Flat-Bottom Quartz Wells
| Fill Volume (µL) | Theoretical Path Length (cm) | KBS-Corrected Path Length (cm) | CV Across 96 Wells (%) |
|---|---|---|---|
| 50 | 0.158 | 0.161 ± 0.004 | 2.5 |
| 100 | 0.315 | 0.320 ± 0.005 | 1.6 |
| 150 | 0.473 | 0.479 ± 0.006 | 1.3 |
| 200 | 0.630 | 0.638 ± 0.007 | 1.1 |
Residual Birefringence and Stress-Induced Optical Artifacts in Quartz
Fused silica manufactured by flame fusion or sol-gel processes retains residual mechanical stress within the glass network unless a controlled annealing cycle reduces the fictive temperature to near equilibrium. Residual stress magnitudes of 0.5–2.5 MPa have been reported in commercially available quartz microplates, corresponding to birefringence1 retardation values of 3–15 nm per centimeter of optical path — measurable with a Sénarmont compensator or a liquid crystal polarimeter.
In standard intensity-based absorbance measurements using unpolarized light, birefringence does not directly alter the measured A260 value because both polarization components are absorbed equally by the chromophore. However, in instruments that use polarized excitation for fluorescence anisotropy — or in UV circular dichroism configurations where quartz plates are occasionally employed — birefringence retardation introduces a systematic polarization rotation that inflates apparent anisotropy values by 5–12 milli-anisotropy units at stress levels above 1.5 MPa.
Inspection for stress birefringence is performed by placing the plate between crossed polarizers under white light illumination; regions exhibiting stress appear as bright interference fringes against the dark field, while stress-free areas remain uniformly dark. Plates showing interference fringes across more than 10% of the well-base area should be rejected for polarization-sensitive measurements, though they remain serviceable for standard intensity-based UV absorbance assays.
Path Length Calibration Method Comparison
| Calibration Method | Required Instrument Feature | Uncertainty (%) | Applicable Solvent |
|---|---|---|---|
| KBS at 977 nm | NIR channel at 977 nm | ±1.5 | Aqueous only |
| KBS at 900 nm | NIR channel at 900 nm | ±4.0 | Aqueous only |
| Geometric calculation | Well diameter measurement | ±5–8 | Any |
| Standard curve back-calculation | Any UV reader | ±2.5 | Any |
| KBS at 977 nm (organic-corrected) | NIR + solvent calibration | ±3.0 | Mixed aqueous/organic |

Reference Standard Selection for 260 nm and 280 nm Assay Calibration
Selecting appropriate reference standards is the penultimate preparatory step before executing the full validation run, and the choice of standard directly determines whether the validated method is traceable to a recognized metrological reference.
Calf thymus DNA (CT-DNA) is the most widely used primary standard for 260 nm calibration due to its well-characterized double-stranded structure, commercially available certified stock solutions, and extinction coefficient of 6,600 L·mol⁻¹·cm⁻¹ per base pair at neutral pH. However, CT-DNA exhibits lot-to-lot variability in GC content (typically 42–45%) that shifts the A260/A230 ratio and can introduce a 3–5% variation in ε₂₆₀ between lots; accordingly, each new lot must be cross-validated against the previous certified lot or against NIST SRM 2366b (Bacillus subtilis genomic DNA), which provides a certified A260 value with a combined measurement uncertainty of ±0.8%. Synthetic oligonucleotide standards with precisely defined sequences offer a higher-precision alternative when the assay target is a defined oligonucleotide, because ε₂₆₀ can be calculated from nearest-neighbor thermodynamics2 to within ±2% and verified by phosphorus elemental analysis.
-
Bovine serum albumin (BSA): The standard reference protein for 280 nm calibration, with ε₂₈₀ = 43,824 M⁻¹·cm⁻¹ for the 66.5 kDa monomer. BSA is appropriate for validating the general photometric performance of the plate reader at 280 nm but does not serve as a proxy for target protein extinction coefficients, which vary by 2–4 orders of magnitude depending on aromatic amino acid composition.
-
IgG reference standards: More relevant than BSA for antibody-focused workflows, with a typical ε₂₈₀ of approximately 210,000 M⁻¹·cm⁻¹ for a 150 kDa IgG1; NIST SRM 927e provides a certified immunoglobulin G concentration standard suitable for traceability documentation.
-
Solvent composition effect on ε values: The extinction coefficients published in literature are universally measured in aqueous buffers at neutral pH. Dissolving nucleic acid or protein standards in TE buffer (10 mM Tris-HCl, 1 mM EDTA, pH 8.0) rather than ultrapure water shifts the A260 baseline by approximately 0.002–0.004 AU due to Tris absorption below 230 nm, which can propagate into the 260 nm measurement if the buffer blank is not rigorously matched. All standard dilutions must be prepared in the same buffer as the experimental blank.
A transition from standard selection to execution requires confirming that standard stocks have been stored according to certified conditions — CT-DNA at −20°C in TE buffer, BSA at 4°C in PBS — and that the stocks have not undergone more than three freeze-thaw cycles, as each cycle introduces approximately 0.5–1.0% degradation in measurable A260 for DNA standards.
Nucleic Acid Validation Runs on Quartz 96 Well Plates
With instrument compatibility confirmed, baseline uniformity established, path length calibrated, and reference standards selected, the full validation run on a quartz 96 well plate proceeds through three principal performance parameters: linearity, precision, and accuracy.
Linearity Range Verification Across Nucleic Acid Concentration Gradients
Linearity is evaluated by preparing a minimum of eight concentration levels spanning the intended working range of the assay, with three replicates per concentration level distributed across non-adjacent wells to decouple positional bias from concentration-dependent effects. For nucleic acid quantification at 260 nm, the recommended concentration range is 0.5–500 ng/µL for dsDNA, covering the full dynamic range of most microplate UV readers at a 100 µL fill volume and a path length of approximately 0.32 cm.
Linear regression of A260 against concentration should yield R² ≥ 0.9990 across the defined range; deviations from linearity at the upper concentration end (typically above 300 ng/µL at 0.32 cm path length) are attributable to the inner filter effect and increased light scattering from condensed chromophores rather than plate or instrument defects. The upper limit of linearity (ULL) is operationally defined as the concentration at which the observed A260 deviates from the predicted regression value by more than 2%; samples above the ULL must be diluted into the linear range before measurement.
At the lower concentration end, the limit of quantification (LOQ) is defined as the concentration producing a signal-to-blank ratio of 10:1, which for a typical quartz microplate reader system at 260 nm corresponds to approximately 0.5–1.0 ng/µL dsDNA in a 100 µL well. This LOQ is approximately 3-fold lower than achievable in polystyrene plates at the same wavelength due to the reduced background absorbance of fused silica, confirming the material-specific advantage of validated quartz 96 well plate systems for trace nucleic acid quantification.
Intra-Assay and Inter-Assay Precision Assessment
Precision is partitioned into two components with distinct experimental designs and acceptance thresholds. Intra-assay precision (repeatability) is measured using a minimum of n = 6 replicate wells loaded with the same concentration of reference standard within a single plate run, with replicates distributed across the plate interior to exclude edge-well positions. The intra-assay CV at 260 nm should not exceed 1.5% for nucleic acid assays and 2.0% for protein assays at 280 nm; CV values above these thresholds at a mid-range standard concentration (e.g., 50 ng/µL dsDNA) indicate residual sources of within-run variability that must be diagnosed and eliminated before the method is considered validated.
Inter-assay precision (intermediate precision) is assessed across a minimum of three independent runs conducted on different days, using freshly prepared standards and a freshly loaded plate for each run. The inter-assay CV acceptance criterion is ≤3.0% for nucleic acid assays and ≤4.0% for protein assays. When inter-assay CV exceeds intra-assay CV by more than a factor of 2.5, the excess variability is typically attributable to day-to-day differences in reagent preparation, analyst pipetting technique, or instrument warm-up state — all of which should be addressed through procedural standardization rather than by relaxing the acceptance criterion.
The experimental design for inter-assay precision must randomize the plate layout across runs so that a given concentration level does not always occupy the same well position in every run; failure to randomize confounds positional bias with run-to-run variability and inflates the apparent inter-assay precision.
Accuracy Verification via Spike-Recovery Experiments
Accuracy is evaluated by the spike-recovery3 method, in which known concentrations of reference standard are added to blank matrix (TE buffer or PBS, as appropriate for the application) at three concentration levels spanning the low, mid, and high regions of the validated linear range. Recovery is calculated as (measured concentration / spiked concentration) × 100%, with an acceptance range of 95–105% for regulated analytical methods and 90–110% for general research applications.
A quartz 96 well plate presents a unique accuracy verification challenge because fused silica surfaces, while chemically inert, exhibit weak electrostatic interactions with positively charged proteins and nucleic acid fragments at near-neutral pH, particularly when the surface has not been pre-blocked or when the buffer ionic strength is below 50 mM. Nucleic acid adsorption onto unblocked fused silica surfaces results in recoveries of 91–94% at concentrations below 5 ng/µL, which falls outside the 95–105% acceptance window and must be addressed by either surface passivation (e.g., brief treatment with 0.1% PEG-silane or BSA pre-coating) or by restricting the validated concentration range to ≥10 ng/µL where adsorption losses are proportionally negligible.
Pre-coating the well surface with 0.1 mg/mL BSA for 15 minutes at room temperature, followed by aspiration and buffer rinse, has been demonstrated to restore DNA recovery to 98.5–102% at concentrations as low as 2 ng/µL, confirming that the accuracy deficit at trace concentrations is surface-chemistry dependent and correctable within the validated protocol.
260/280 Ratio Fidelity as a Purity Indicator
The A260/A280 ratio is the primary spectrophotometric purity indicator for nucleic acid preparations, with accepted reference values of 1.80 ± 0.05 for purified dsDNA and 2.00 ± 0.05 for purified RNA in TE buffer at pH 8.0. Validation of ratio fidelity requires demonstrating that the plate reader — operating in conjunction with the quartz 96 well plate — reproduces the accepted reference ratios for certified reference standards within the stated tolerance, without systematic bias.
Ratio bias most commonly arises from wavelength inaccuracy rather than photometric linearity errors; a −0.5 nm offset at the 260 nm setpoint of a monochromator-based reader reduces the apparent A260 by approximately 1.5%, shifting the A260/A280 ratio of a pure dsDNA sample from 1.80 to approximately 1.77 — a deviation that would incorrectly suggest protein contamination in an otherwise pure preparation. For this reason, ratio fidelity validation must be coupled with the wavelength accuracy verification described in the instrument compatibility chapter, and any identified wavelength offset must be corrected before ratio-dependent purity assessment is performed.
Plate-material-specific contributions to ratio bias are generally minimal in high-quality fused silica plates, as the plate absorbance at 280 nm is typically <0.005 AU lower than at 260 nm for identically thick well bases — a differential small enough to be fully accounted for by the blank subtraction procedure without introducing ratio bias above ±0.01.
Validation Run Performance Parameters
| Performance Parameter | Nucleic Acid (260 nm) | Protein (280 nm) | Acceptance Limit |
|---|---|---|---|
| Linearity R² | ≥0.9990 | ≥0.9985 | Per application |
| Dynamic range (ng/µL) | 0.5–500 | 5–2000 | Assay-dependent |
| LOQ (ng/µL) | ~0.5–1.0 | ~5 | S/N ≥ 10 |
| Intra-assay CV (%) | ≤1.5 | ≤2.0 | Single-run, n ≥ 6 |
| Inter-assay CV (%) | ≤3.0 | ≤4.0 | 3 days, n = 3 runs |
| Spike recovery (%) | 95–105 | 95–105 | Mid-range standard |
| A260/A280 ratio bias | ≤±0.03 | N/A | Vs. cuvette reference |

Protein Quantification Validation at 280 nm with Quartz Microplates
Direct UV absorbance at 280 nm offers a reagent-free, rapid protein quantification method whose accuracy depends critically on the extinction coefficient used for calculation and the correction strategy applied to turbid or contaminated samples.
Extinction Coefficient Selection for Target Proteins
The molar extinction coefficient at 280 nm (ε₂₈₀) for a given protein is determined primarily by its content of tryptophan (ε = 5,500 M⁻¹·cm⁻¹ per Trp) and tyrosine (ε = 1,490 M⁻¹·cm⁻¹ per Tyr) residues, with a minor contribution from disulfide bonds (ε ≈ 125 M⁻¹·cm⁻¹ per S–S bond). Proteins with zero tryptophan residues — such as parvalbumin or some short peptide hormones — exhibit ε₂₈₀ values below 500 M⁻¹·cm⁻¹, making direct A280 quantification impractical at concentrations below 1 mg/mL in a microplate format with a 0.32 cm path length.
Using BSA (ε₂₈₀ = 43,824 M⁻¹·cm⁻¹) as a universal surrogate extinction coefficient for any target protein introduces concentration errors proportional to the difference in aromatic content between BSA and the target. For an antibody with ε₂₈₀ = 210,000 M⁻¹·cm⁻¹, applying the BSA coefficient would underestimate concentration by approximately 4.8-fold. Accurate quantification requires using the sequence-specific theoretical ε₂₈₀ calculated by the ExPASy ProtParam tool from the UniProt-deposited amino acid sequence, or the experimentally determined ε₂₈₀ measured by quantitative amino acid analysis. The ExPASy-calculated ε₂₈₀ agrees with experimentally determined values within ±5% for the majority of soluble globular proteins, with larger deviations observed for membrane proteins with unusual aromatic distributions.
When the target protein sequence is proprietary or unavailable, a conservative approach is to calculate an empirical ε₂₈₀ by measuring the A280 of a sample whose concentration has been independently determined by a non-UV method (such as bicinchoninic acid assay or amino acid analysis) and back-calculating ε₂₈₀ from Beer-Lambert. This empirically derived coefficient must be documented as a method-specific parameter in the validation record.
Scattering Correction for Turbid Protein Samples in Quartz Wells
Protein samples containing aggregates, lipid particles, or colloidal contaminants scatter incident light in a wavelength-dependent manner described approximately by Rayleigh scattering (I_scatter ∝ λ⁻⁴), which produces a rising absorbance baseline as wavelength decreases from 350 nm toward 260 nm. In turbid protein samples measured in a quartz microplate, the apparent A280 can be inflated by 0.02–0.15 AU depending on the aggregate concentration — a systematic positive error that would cause protein concentration to be overestimated by 10–50% if uncorrected.
The standard correction method involves measuring absorbance at a reference wavelength in the UV-transparent window where no protein chromophore absorbs, typically 320 nm or 340 nm, and subtracting the scattering contribution at 280 nm using the wavelength-dependence of Rayleigh scattering: A₂₈₀_corrected = A₂₈₀_measured − A₃₂₀ × (320/280)⁴. Applied consistently, this correction reduces scattering-attributable error to below ±3% for samples with A₃₂₀ values up to approximately 0.05 AU.
Fused silica's intrinsically low autofluorescence and UV background absorbance — typically <0.003 AU at 320 nm for a clean plate — make it significantly more reliable than UV-transparent polystyrene for scattering correction, because polystyrene plates exhibit a measurable absorbance slope between 300 and 340 nm that confounds the scattering baseline measurement. This material-specific advantage of the quartz 96 well plate format is particularly valuable in workflows involving cell lysates, crude protein extracts, or lipid nanoparticle formulations where turbidity is inherent to the sample matrix.
Protein Quantification Validation Parameters
| Parameter | Value / Criterion | Method |
|---|---|---|
| Tryptophan ε₂₈₀ (M⁻¹·cm⁻¹) | 5,500 per residue | Edelhoch method |
| Tyrosine ε₂₈₀ (M⁻¹·cm⁻¹) | 1,490 per residue | Edelhoch method |
| ExPASy ε₂₈₀ accuracy (%) | ±5 vs. experimental | Globular proteins |
| Scattering correction wavelength (nm) | 320 or 340 | Rayleigh model |
| Scattering A₃₂₀ upper limit (AU) | ≤0.05 | Before correction |
| Post-correction accuracy (%) | ±3 | A₃₂₀ ≤ 0.05 AU |
| Plate background at 320 nm (AU) | <0.003 | Clean fused silica |
Cleaning, Regeneration and Reuse Qualification of Quartz Well Plates
Given the substantial material cost of fused silica microplates, reuse qualification is a practical necessity — and cleaning efficacy must be validated with the same rigor applied to the original performance characterization.
-
Protein and nucleic acid contaminants: Hellmanex III at 1% (v/v) in ultrapure water at 60°C for 30 minutes effectively removes adsorbed proteins and DNA from fused silica surfaces, with residual protein quantified by BCA assay on the rinse fraction typically falling below 0.5 ng/cm² after a single treatment cycle. A final rinse with ultrapure water (3× by volume) is required to remove detergent residues that would otherwise absorb at 260 nm and inflate blank readings by up to 0.008 AU.
This cleaning approach is supported by surface contact angle measurements showing recovery to <5° water contact angle after Hellmanex treatment, consistent with a hydroxyl-terminated, contamination-free fused silica surface. Verification of cleaning efficacy should include a post-cleaning blank absorbance check confirming that A260 returns to within ±0.003 AU of the pre-use validated baseline.
-
Fluorescent dye residues: Intercalating dyes (SYBR Green, ethidium bromide) and protein-reactive fluorophores (Alexa Fluor series) require more aggressive removal; 10% (v/v) sodium hydroxide at room temperature for 20 minutes followed by thorough rinsing is effective for anionic dyes. UV/ozone treatment (254 nm, 15 minutes) provides a complementary non-chemical decontamination for dyes resistant to alkaline hydrolysis, reducing fluorescence background by >95% as measured by plate reader scan at the dye's excitation wavelength.
A transition point in the cleaning protocol occurs after NaOH treatment: the elevated pH must be fully neutralized before UV absorbance re-verification, as residual alkalinity shifts the apparent A260 of any water blank through increased UV absorption by hydroxide ions above pH 10.
-
Reuse qualification and retirement criteria: After each cleaning cycle, inter-well blank CV and edge-to-center gradient are re-measured and compared against the original validation baseline. A plate is retired from validated use when the inter-well CV at 260 nm exceeds 2.5% on two consecutive post-cleaning qualification runs, or when any individual well exhibits a persistent blank A260 deviation >0.010 AU from the plate mean despite repeated cleaning, indicating irreversible surface modification. Empirical observations from multi-cycle reuse studies show that fused silica plates subjected to NaOH cleaning maintain acceptable performance for 15–25 cleaning cycles before CV degradation reaches the retirement threshold, while plates cleaned exclusively with Hellmanex retain acceptable performance for 30–50 cycles under typical laboratory conditions.
Documentation and Traceability Requirements for UV Assay Validation Records
For laboratories operating under GMP, GLP, or ISO 17025 quality frameworks, the technical validity of a UV assay validation is inseparable from the completeness and integrity of its associated documentation.
-
Core validation report elements: Every validation record for a quartz 96 well plate UV assay must include the plate manufacturer, catalog number, and production lot number; the plate reader serial number, firmware version, and most recent calibration certificate date; the reference standard certificate of analysis with certified concentration and traceability to NIST or equivalent national metrology institute; all raw absorbance data files in non-editable format; the identity, date, and organizational affiliation of the analyst performing each validation run. Omission of any of these elements creates a traceability gap that invalidates the document for regulatory submission purposes.
Within the validation report, each performance parameter (linearity, precision, accuracy, ratio fidelity) should be presented in a table format alongside its acceptance criterion, the observed value, and a pass/fail designation — structured to enable rapid auditor review without reference to the underlying raw data files.
-
21 CFR Part 11 electronic record compliance: Laboratories within FDA-regulated environments that capture validation data in electronic laboratory notebooks (ELNs) or plate reader software must ensure that data files are stored in audit-trail-enabled, time-stamped formats that prevent post-acquisition modification without a traceable record. Plate reader software compliant with 21 CFR Part 11 — such as Tecan i-control with FDA module or Molecular Devices SoftMax Pro GxP — generates electronic signatures linked to individual user credentials, satisfying the identity verification requirement of the regulation. Raw data exported to Excel or CSV formats loses audit trail integrity and is not considered compliant without supplementary procedural controls.
-
Plate use logbooks: Each individual quartz microplate should be assigned a unique identifier (either manufacturer serial number or laboratory-assigned barcode) and tracked in a use logbook recording the date of each use, the assay performed, the analyst, the cleaning method applied, and the post-cleaning qualification result. This logbook enables retrospective identification of any validation run performed on a plate that subsequently failed qualification, allowing affected data to be flagged for review. The logbook also provides the empirical basis for establishing plate-specific retirement timelines, replacing generic manufacturer recommendations with data derived from the plate's actual use history under the laboratory's specific conditions.
Conclusion
Validating a quartz 96 well plate for UV absorbance at 260 and 280 nm requires addressing six distinct technical domains in sequence: substrate optical characterization, instrument compatibility, baseline uniformity, path length calibration, analytical performance verification, and documentation. Each domain contains specific, quantifiable acceptance criteria — from inter-well CV thresholds of ≤2.0% to spike recovery ranges of 95–105% — that collectively define a validated, defensible measurement system. Laboratories that execute this protocol in full obtain not only regulatory-compliant data but also a quantitative understanding of every error source in their UV quantification workflow, enabling confident interpretation of nucleic acid purity and protein concentration results across the full dynamic range of the fused silica microplate format.
FAQ
What fill volume gives the most accurate path length correction in a quartz 96 well plate?
A fill volume of 150–200 µL provides the most accurate path length correction in a standard flat-bottom quartz 96 well plate, because the larger liquid column height reduces the proportional contribution of meniscus geometry and well-bottom thickness variation to total path length uncertainty. At 200 µL, the KBS-corrected path length CV across 96 wells typically falls to 1.1%, compared to 2.5% at 50 µL.
Can a quartz 96 well plate be used without path length correction if only ratio data is needed?
A260/A280 ratio measurements are relatively insensitive to absolute path length errors because both wavelengths experience the same path, and the ratio cancels out most multiplicative path-length factors. However, wavelength-dependent path length variation — arising from chromatic aberration in the collection optics or refractive index dispersion in the fused silica — introduces a small wavelength-dependent path difference that can shift the A260/A280 ratio by ±0.02–0.04 at sub-optimal fill volumes. Path length correction is recommended even for ratio-only applications when working below 50 µL.
How many reuse cycles can a quartz microplate sustain before UV performance degrades?
Under Hellmanex III cleaning at 1% concentration, quartz 96 well plates typically sustain 30–50 cleaning cycles before inter-well CV at 260 nm exceeds the 2.5% retirement threshold. Plates cleaned with 10% NaOH show earlier surface hydroxylation changes and typically reach the retirement threshold after 15–25 cycles. Individual plate performance varies with the severity of contaminants processed, and periodic re-qualification after every 10 cycles is recommended regardless of cleaning method.
Does buffer composition affect blank absorbance in fused silica microplates at 260 nm?
Tris-HCl buffer at concentrations above 20 mM absorbs measurably below 230 nm, and at 10 mM contributes approximately 0.001–0.002 AU at 260 nm — negligible in most applications but relevant for samples near the LOQ. EDTA at 1 mM contributes <0.001 AU at 260 nm and is non-interfering. PBS (phosphate-buffered saline) is spectrally transparent at 260 nm and 280 nm and is the preferred blank buffer when matrix matching between blank and sample is not feasible with TE buffer.
References:
-
Birefringence is the optical property of a material in which the refractive index differs along different crystallographic or stress axes, causing incident light to split into two polarized components that travel at different speeds through the medium. ↩
-
Nearest-neighbor thermodynamics is a computational model used to predict the thermodynamic stability and molar extinction coefficients of oligonucleotide sequences based on the stacking interactions between adjacent base pairs, enabling precise ε₂₆₀ calculation for synthetic DNA and RNA standards. ↩
-
Spike-recovery is an analytical accuracy assessment method in which a known amount of reference analyte is added to a sample matrix, and the percentage of that analyte subsequently measured by the method is calculated to evaluate matrix effects and systematic bias. ↩